If you are working with pipes in R, check out my top 10 favorite dplyr tips and tricks. 
In this tutorial, let’s explore how to search multiple strings using only one grep process. Snames %>% grepl(paste(p, collapse = "|"). Overview When we work in the Linux command line, we often use the grep command to search text. snames % stringr::str_detect(paste(p, collapse = "|")) With two strings, it is not a lot of work but below is a more efficient approach that will be useful if there is a lot of text patterns. I know that there is a -o flag that can be used to get only matches. It finds matches correctly, but I can't figure out how to output all lines in a separate file, excluding matches within them. If you want to find out the appearance of multiple strings at once, then first and the simplest approach is a combination of multiple results of grepl. I use the following command to search for such strings: grep -P ' (<>) A-Z0-9\.\s' File.txt.Detect combination of multiple strings at once in R There will be two main scenarios – detection of multiple strings simultaneously and any of the given strings. Note that even if you typed data as input to the first grep, you would only get any lines that contain 'word2' shown on the output.Īt one time, the answer used: grep -E 'word1.*word2|word2.*word1' 'word1.*word2' triggered the comments below.Here is how to detect multiple strings in R using base function grepl and alternatives, like str_detect from package stringr. Now you have two processes running, the first grep and the shell, and they are both trying to read from the keyboard, and it is not determinate which one gets any given line of input (or any given EOF indication). With Heirloom Shell, the second grep completes and exits, and the shell prompts again. With Bash, the parent shell waits until all the processes in the pipeline have exited, so it sits around waiting for the grep -c to finish, which it won't do until you indicate EOF. Note that the second grep doesn't read its standard input at all, so the pipe is superfluous. Or, in my case, it will fail since there's neither a file nor a directory called logs where I'm running the pipeline. The second grep does a recursive search for 'word2' in the files underneath directory logs (or, if it is a file, in the file logs). by typing Control-D), it will sit there, waiting for you to type something. type f xarg grep -il osuser but i have one more string 'vsession' here i want to grep in which file these two strings are present. The first grep is going to count the number of occurrences of 'word1' on its standard input, and print that number on its standard output. 1 01-19-2011 gagan4599 Registered User 13, 0 Grep multiple strings in multiple files using single command Hi, I will use below command for grep single string ('osuser' is search string) ex: Code: find. The initial script is: grep -c "word1" | grep -r "word2" logs If you run two commands, you should aim to select the less frequently occurring word in the first grep to minimize the amount of data processed by the second. Against that, the regex is more complex you might need to benchmark it to find out what's best - but only if performance really matters. However, if the file is huge and 'word1' occurs frequently, then you may be passing significant data down the pipe where a single command avoids that overhead. If the file is big but only a few lines contain 'word1', there isn't going to be much data passed on the pipe and using two command is fine. If the file is small, performance isn't likely to be an issue and running two commands is fine. How much this matters depends on how big the data file is and how many lines match 'word1'. In this article, we’re going to show you how to use GNU grep to search for multiple strings or patterns. The 'one command' variants have the advantage that there is only one process running, and so the lines containing 'word1' do not have to be passed via a pipe to the second process. grep is a powerful command-line tool that allows you to searches one or more input files for lines that match a regular expression and writes each matching line to standard output. If you know that 'word1' will precede 'word2' on the line, you don't even need the alternatives and regular grep would do: grep 'word1.*word2' logs 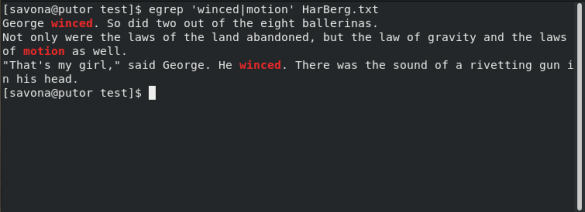
You could use extended grep ( grep -E or egrep): grep -E 'word1.*word2|word2.*word1' logs However, it isn't necessary to use two commands like that. The first grep finds lines with 'word1' from the file 'logs' and then feeds those into the second grep which looks for lines containing 'word2'. One simple rewrite of the command in the question is: grep "word1" logs | grep "word2" 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
